Transfers attribute data from a source spatial layer to a target spatial layer based
on the area of overlap between their geometries. All calculations are performed in
SedonaDB for efficiency.
Supports lazy evaluation returning sedonadb_dataframe objects.
Usage
sx_interpolate_aw(
target,
source,
tid,
sid,
extensive = NULL,
intensive = NULL,
weight = "sum",
output = NULL,
view_name = NULL,
keep_NA = TRUE,
na.rm = FALSE,
join_crs = NULL,
verbosity = NULL,
use_s2 = NULL,
...
)Arguments
- target
A
sedonadb_dataframe,sfobject, or view name (character) in SedonaDB representing destination geometries.- source
A
sedonadb_dataframe,sfobject, or view name (character) in SedonaDB containing data to interpolate.- tid
Character. Unique ID column name in
target.- sid
Character. Unique ID column name in
source.- extensive
Character vector. Columns in
sourceto be treated as extensive (counts).- intensive
Character vector. Columns in
sourceto be treated as intensive (rates).- weight
Character. Denominator for extensive variables: "sum" (default) or "total".
- output
Character or NULL. Output type:
sedonadb_dataframe(default),sf,tibble,geoarrow, orraw. If NULL, usesgetOption("sx.output_type", "sedonadb_dataframe").Output types:
sedonadb_dataframe: Lazy data frame (no collection).sf: Materialized sf object.tibble: Tibble without geometry.geoarrow: Tibble withgeoarrow_vctrgeometry (Arrow-native).raw: Tibble with geometry as raw WKB bytes (for database import).
- view_name
Character (optional). Name to register the result as a persistent view in the active backend. If NULL (default), returns the result directly without creating a view.
Not all backends support named views. Check backend-specific documentation for availability.
- keep_NA
Logical. If TRUE, output includes all target features (LEFT JOIN).
- na.rm
Logical. If TRUE, source features with NA values are ignored.
- join_crs
Numeric or Character (optional). EPSG code or WKT for CRS transform during calc.
- verbosity
Character or NULL. Controls message output for this function call.
"quiet": Suppress all informational messages."info": Show standard progress and status messages."debug": Show additional diagnostic messages for troubleshooting.
If NULL (the default), uses the global
sx.verbosityoption. Seesx_options()for persistent configuration.- use_s2
Logical or NULL. Controls spherical geometry (S2) for this operation.
TRUE: Use S2 spherical geometry (accurate for geographic coordinates).FALSE: Use planar geometry (faster, appropriate for projected CRS).NULL(default): Uses the globalsx_use_s2()setting.
- ...
Ignored. Used to catch and warn about unsupported sf arguments.
Details
Areal-weighted interpolation assumes uniform distribution of values within source polygons.
Coordinate Systems:
Area calculations are sensitive to CRS. It is strongly recommended to use a projected CRS.
Use the join_crs argument to project data on-the-fly during the interpolation.
Extensive vs. Intensive Variables:
Extensive (counts, sums): Value is divided proportionally to area. Use
weight="sum"(relative to target coverage) orweight="total"(relative to source area).Intensive (rates, densities): Value is averaged based on partial areas. Always uses intersection area weighting.
See also
areal::aw_interpolate() for reference implementation.
Examples
# \donttest{
library(sf)
# 1. Prepare Data
# Load NC counties (source) and project to Albers (EPSG:5070)
nc <- st_read(system.file("shape/nc.shp", package = "sf"), quiet = TRUE)
nc <- st_transform(nc, 5070)
nc$sid <- seq_len(nrow(nc))
# Create a target grid
grid <- st_make_grid(nc, n = c(10, 5)) |> st_as_sf()
grid$tid <- seq_len(nrow(grid))
# -------------------------------------------------------------------
# Example 1: Using sf objects directly (most common use case)
# -------------------------------------------------------------------
# Extensive interpolation (total counts, e.g., births)
result_ext <- sx_interpolate_aw(
target = grid, source = nc,
tid = "tid", sid = "sid",
extensive = "BIR74",
weight = "total",
output = "sf"
)
# Check mass preservation (should be ~1.0)
sum(result_ext$BIR74, na.rm = TRUE) / sum(nc$BIR74)
#> [1] 1
# Intensive interpolation (rates/densities)
result_int <- sx_interpolate_aw(
target = grid, source = nc,
tid = "tid", sid = "sid",
intensive = "BIR74",
output = "sf"
)
# -------------------------------------------------------------------
# Example 2: Using sedonadb_dataframe (lazy evaluation)
# -------------------------------------------------------------------
# First operation returns lazy result
lazy_result <- sx_interpolate_aw(
target = grid, source = nc,
tid = "tid", sid = "sid",
extensive = c("BIR74", "BIR79"),
output = "sedonadb_dataframe"
)
# Materialize when ready
final_sf <- sx_collect(lazy_result)
# -------------------------------------------------------------------
# Example 3: Using pre-registered SedonaDB view names
# -------------------------------------------------------------------
# Register data as views
sx_as_view(nc, "nc_counties")
sx_as_view(grid, "target_grid")
# Use view names as input
result_from_views <- sx_interpolate_aw(
target = "target_grid", source = "nc_counties",
tid = "tid", sid = "sid",
extensive = "BIR74",
output = "sf"
)
# Quick visualization
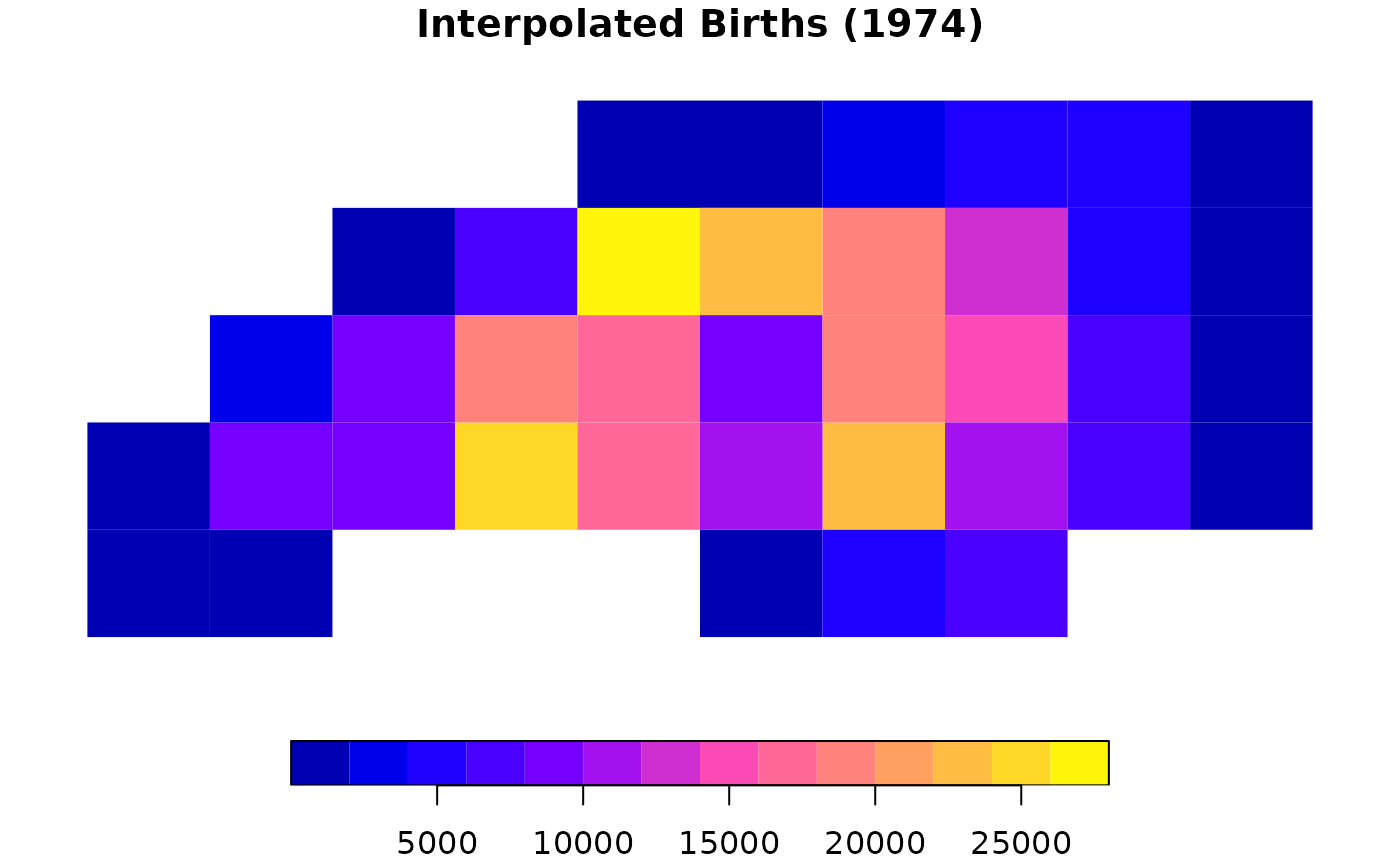
plot(result_ext["BIR74"], main = "Interpolated Births (1974)", border = NA)
 # -------------------------------------------------------------------
# Example 4: Arrow ecosystem
# -------------------------------------------------------------------
# Export as geoarrow for zero-copy Parquet writing
geo_result <- sx_interpolate_aw(grid, nc, "tid", "sid", extensive = "BIR74", output = "geoarrow")
# }
# -------------------------------------------------------------------
# Example 4: Arrow ecosystem
# -------------------------------------------------------------------
# Export as geoarrow for zero-copy Parquet writing
geo_result <- sx_interpolate_aw(grid, nc, "tid", "sid", extensive = "BIR74", output = "geoarrow")
# }
